Mpox surveillance in Congo identifies three viral clades circulating at once
A newly published report in
Nature Medicine
adds an important layer to the global mpox picture: researchers say three distinct monkeypox virus clades and lineages were found circulating in the Republic of the Congo in 2025. The study describes a laboratory-confirmed case of mpox clade IIb lineage A.2.2 in Pointe-Noire, the country’s second-largest city, and places that finding alongside additional cases from clade Ia, clade Ib, and another introduction of clade IIb detected through passive surveillance.The headline matters because clade IIb is the branch associated with the 2022 global epidemic driven largely by human-to-human transmission. According to the paper, published data on circulation of clade IIb in central Africa had been absent before this report. The new evidence therefore does more than log a single case. It suggests that the regional epidemiological landscape is more complicated than a single outbreak narrative and that health systems may be confronting several overlapping transmission patterns at once.
What the researchers found
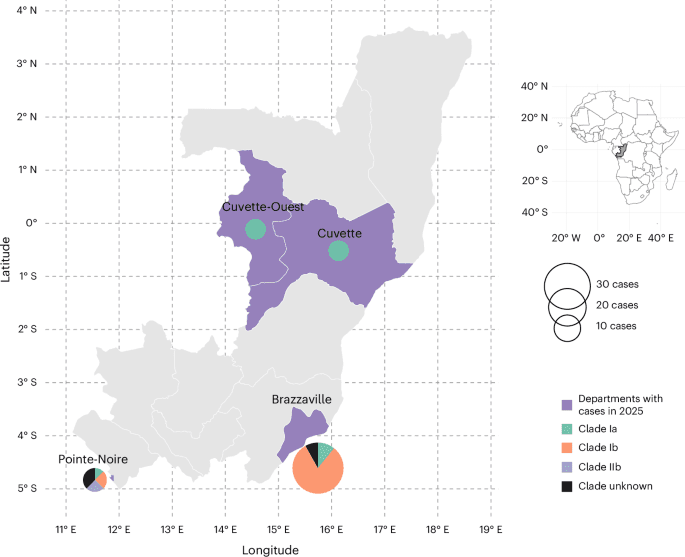
The authors report one confirmed case of mpox clade IIb lineage A.2.2 in Pointe-Noire. Whole-genome phylogenetic analysis placed that virus within a lineage now emerging in West Africa, particularly Sierra Leone. Beyond that case, the study says passive surveillance in the Republic of the Congo identified 16 cases of clade Ia, 32 cases of clade Ib, and one additional introduction of clade IIb during 2025.
That combination, the researchers write, marks the co-circulation of a third distinct mpox clade and lineage in the human population, alongside clades Ia and Ib. In practical terms, that means public health teams cannot assume that one detected cluster fully represents what is moving through communities. Diagnostic, surveillance, and response systems need to be able to distinguish among clades and lineages rather than treating all mpox detections as interchangeable.
Why the finding matters
The study’s strongest message is operational. The authors argue that improved surveillance and diagnostic strategies are needed to identify which clade and lineage are circulating in the human population. That is a significant point because public health decision-making depends on timely, accurate characterization of outbreaks. A more diverse viral picture raises the stakes for laboratory capacity, genomic sequencing, and case tracking.
The paper also ties the Congo findings to broader risk. The researchers call for stronger regional capacity for case detection, contact tracing, and public health measures, along with affordable vaccines. Their warning is explicit: without better preparedness and response, both clade I and clade II mpox continue to pose a wider global threat.
For policymakers and outbreak monitors, this report is a reminder that mpox is not a static post-2022 story. The virus continues to evolve in geographic reach and lineage complexity, and surveillance gaps can obscure that reality until genomic evidence catches up. For scientists, the Congo data add a new reference point in tracking how different branches of mpox are moving within and across regions.
The immediate takeaway is not that all transmission patterns are the same, but that public health systems need the tools to tell the differences apart. This study makes the case that better genomic and diagnostic visibility is no longer optional.
This article is based on reporting by Nature Medicine. Read the original article.