Seeing What Standard Tests Miss
New research demonstrates that optical genome mapping (OGM) can detect clinically significant genetic variants that conventional diagnostic methods miss in nearly one out of every five patients with acute leukemia. The findings, which could alter treatment decisions for a substantial fraction of leukemia patients, add to growing evidence that OGM technology is ready to complement or replace some traditional cytogenetic testing methods in clinical oncology.
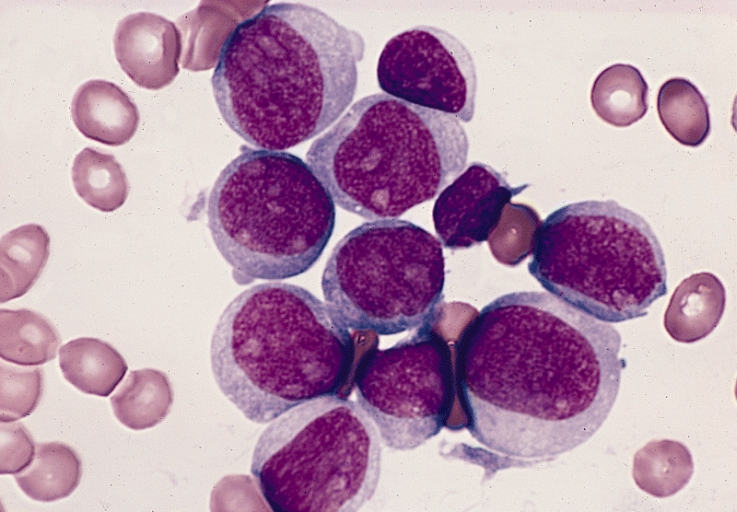
Acute leukemia — which includes acute myeloid leukemia (AML) and acute lymphoblastic leukemia (ALL) — is driven by genetic mutations that cause blood cell precursors to multiply uncontrollably. Identifying the specific mutations present in a patient's leukemia is critical for treatment planning, as different genetic subtypes respond to different therapies.
How Optical Genome Mapping Works
Traditional cytogenetic methods for detecting genetic abnormalities in leukemia include karyotyping — visually examining chromosomes under a microscope — and fluorescence in situ hybridization (FISH), which uses fluorescent probes to detect specific chromosomal regions. These methods have been standard practice for decades but have significant limitations.
Karyotyping requires cells to be cultured and dividing, which fails in a significant fraction of leukemia samples. FISH can only detect the specific genetic targets it is designed to probe. Both methods have limited resolution for detecting smaller structural variants.
Optical genome mapping takes a fundamentally different approach. Instead of examining individual cells, OGM extracts ultra-long DNA molecules from patient samples, labels them with fluorescent tags at specific sequence motifs, and passes them through nanochannels where high-resolution cameras image the fluorescent pattern. Software then assembles these individual molecule images into a genome-wide map, comparing it against a reference genome to identify structural variants.
The key advantage is the ability to detect structural variants across the entire genome in a single assay, without the need for cell culture or prior knowledge of which variants to look for.