A new H5N1 genotype surged across migratory flyways
A study published online in Nature Medicine on April 15, 2026 reports that a newly classified high-pathogenicity avian influenza genotype, D1.1, expanded rapidly in wild birds across North America during the 2024 migratory season. The paper describes the virus as a reassortant that was first detected in September 2024 and then tracked through active and passive surveillance programs in Canada and the United States.
The central finding is not merely that H5N1 remained present in wild bird populations, but that a distinct genotype appears to have spread quickly enough to displace earlier A(H5) lineages across several flyways. That matters because migratory flyways are the channels through which avian influenza can move over long distances, cross jurisdictions, and repeatedly reseed outbreaks in new places.
By tying genomic surveillance to the seasonal movement of wild birds, the study adds a sharper picture of how a specific viral lineage can go from emergence to broad geographic reach in a short period. It also shows how much depends on maintaining surveillance systems that can detect those shifts before they become visible in livestock or human case counts.
What the researchers reported
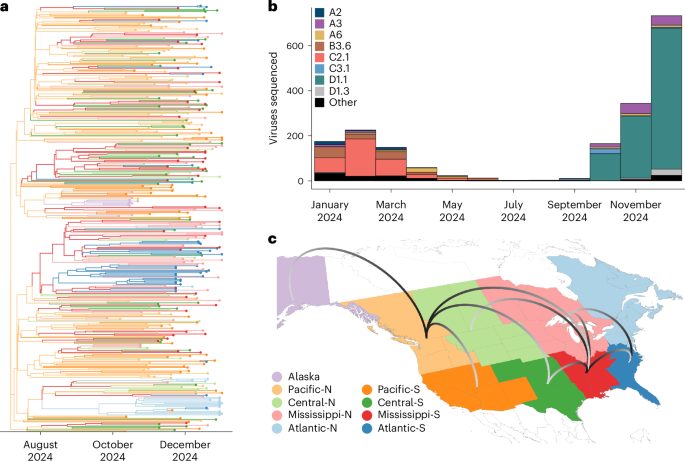
According to the abstract supplied with the candidate, highly pathogenic avian influenza A(H5N1) clade 2.3.4.4b viruses entered North America in late 2021 and then reassorted rapidly with local avian influenza viruses. The newly observed D1.1 genotype was detected in September 2024. Using surveillance data from across Canada and the US, the researchers followed its emergence and spread during the fall migration.
The study says phylodynamic analysis showed that D1.1 viruses formed a monophyletic group. In practical terms, that supports the idea that the viruses tracked in the surveillance network belonged to a coherent, newly expanded lineage rather than a loose collection of unrelated detections. The paper further states that D1.1 displaced earlier A(H5) genotypes across several flyways, underscoring that this was not a marginal event at the edges of the surveillance map.
The source text also links the expansion of D1.1 with detections in other hosts, including 17 human cases, four of which were severe or fatal. At the same time, the abstract notes that the mammalian-adaptive markers found in human cases were not detected in the wild-bird viruses analyzed in the study. That distinction is important: it suggests the surveillance findings in wild birds did not directly show the same adaptive signatures reported in the human cases.
Why this matters beyond bird surveillance
The study lands at the intersection of wildlife ecology, animal health, and human health. A rapidly expanding genotype in wild birds is not only a conservation or veterinary issue; it is also a warning about the pace at which influenza ecology can change. When a lineage becomes established across migratory routes, the number of opportunities for spillover events, agricultural exposure, and cross-species transmission rises.
The supplied abstract stops short of claiming that D1.1 itself acquired the mammalian-adaptive markers seen in human infections, and that caution is part of the point. Influenza risk is shaped by a moving combination of viral genetics, host exposure, and ecological opportunity. A genotype can become epidemiologically important even without immediately showing every mutation associated with adaptation in people.
That makes early, broad genomic surveillance especially valuable. The paper’s use of both active and passive surveillance signals that no single collection method is enough for a virus that can circulate across a continent. Detecting a new genotype is one step. Understanding whether it is replacing others, how widely it is moving, and whether it is appearing in additional hosts are separate questions that require sustained sampling.
What the paper does and does not say
The source material supports several clear conclusions. D1.1 emerged as a newly detected reassortant in September 2024. It spread rapidly in wild birds during the 2024 fall migration. It formed a monophyletic group in the authors’ analysis and displaced earlier A(H5) genotypes across several flyways. Candidate vaccine viruses, the abstract adds, retained antigenic cross-reactivity with D1.1 strains.
That last point is notable because it suggests that, based on the authors’ reported findings, vaccine candidates were not rendered antigenically irrelevant by the rise of this genotype. It does not end the risk discussion, but it does indicate that viral change and vaccine preparedness did not immediately diverge in the way public-health officials most fear.
The abstract does not provide a full geographic breakdown, a month-by-month chronology of displacement, or the complete details of the human case context. Those details may be present in the full paper, but they are not contained in the supplied source text. What can be said with confidence is that the study documents a fast and consequential reshaping of the H5N1 landscape in North American wild birds during one migration season.
A signal for the surveillance era
The broader lesson is that influenza surveillance is increasingly a race against viral recombination and movement. By the time a lineage is widely discussed in public, it may already have traversed multiple flyways and crossed into multiple host settings. The D1.1 report shows why genomic tracking has become essential infrastructure rather than a niche research tool.
For policymakers and health agencies, the study reinforces a familiar but still urgent message: emerging influenza threats are often first visible in ecological systems, not hospitals. For researchers, it offers a case study in how quickly a reassortant lineage can establish itself. And for the wider public, it is a reminder that avian influenza stories are no longer isolated farm or wildlife incidents. They are continental systems events that require sustained attention long before any headline count of human cases arrives.
This article is based on reporting by Nature Medicine. Read the original article.
Originally published on nature.com